Oligosaccharyltransferase Complex
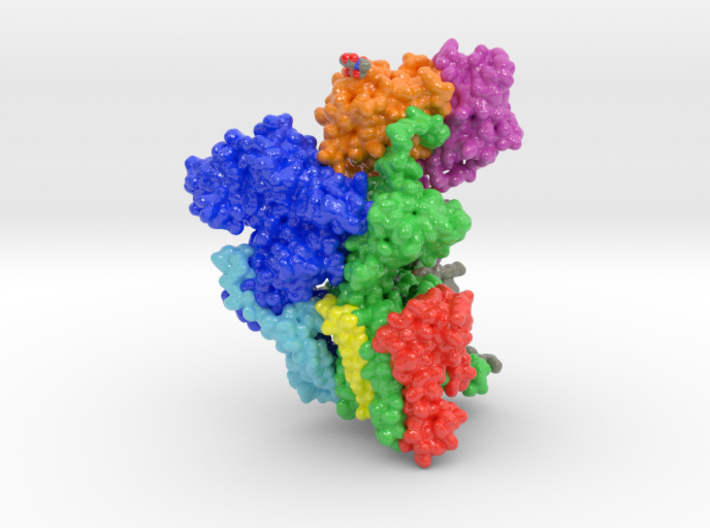
This is a 3D print of oligosaccharyltransferase (OST) complex. It is colored blue according to the protein’s electrostatic property. Colored green are various monosaccharides attached to OST as sugar trees. Isolated and colored by atom type are the binding residues that attach shuttling proteins. Colored white are individual phospholipids from a plasma membrane.
Protein Description 6EZN
When cells interact with bacteria, viruses or other cells, a complex exchange of protein interactions occurs. Sometimes, membrane-bound proteins internalize based these interactions with external stimuli. When this occurs, sugar trees attached to these exterior proteins also internalize. When sugar aggregates, they form tree-like structures called glycans that attach to membrane lipids or proteins with available binding residues. Sugar is both necessary for cellular respiration and ATP production, but also dangerous causing cellular dysfunction if exposed to large quantities. Understanding how sugar is shuttled into cells is an important topic of study.
Biologic Explorer: 6EZN
Http iframes are not shown in https pages in many major browsers. Please read this post for details.Oligosaccharyltransferase (OST) is an essential membrane protein complex in the Endoplasmic Reticulum, where it transfers an oligosaccharide from a dolichol-pyrophosphate-activated donor to glycosylation sites of secretory proteins. We here describe the atomic structure of yeast OST determined by cryo-EM, revealing a conserved subunit arrangement. The active site of the catalytic STT3 subunit points away from the center of the complex, allowing unhindered access to substrates. The dolichol-pyrophosphate moiety binds to a lipid-exposed groove of STT3, while two non-catalytic subunits and an ordered N-glycan form a membrane-proximal pocket for the oligosaccharide. The acceptor polypeptide site faces an oxidoreductase domain in standalone OST complexes or is immediately adjacent to the translocon, suggesting how eukaryotic OSTs efficiently glycosylate a large number of polypeptides prior to their folding.







