Description
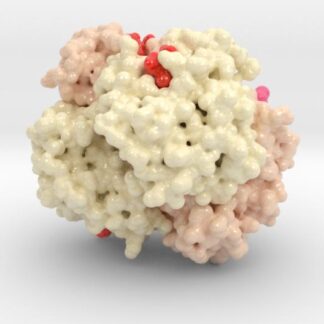
Hemoglobin HbS tetramers have been shown to be the fundamental building block of the intracellular sickle cell fiber causing the effects of Sickle Cell Disease.
Model Description
Biologic model of Sickle Cell Hemoglobin HbS 3D printed in full-color sandstone. and visualizes the HbS mutation locations (pink).
For international customers outside the US, please visit the model on our 3D printing service’s international website: LINK
Select the desired material finish and size below. Matte finish applies a UV protective, semi-glossy coating. Natural finish does not.
Created from PDB ID: 2HBS